Tag: drug discovery

A smarter way to streamline drug discovery
The use of AI to streamline drug discovery is exploding. Researchers are deploying machine-learning models to help them identify molecules, among billions of options, that might have the properties they are seeking to develop new medicines.But there are so many variables to consider — from the price of materials to the risk of something going wrong — that even when scientists use AI, weighing the costs of synthesizing the best candidates is no easy task.
The myriad challenges involved in identifying the best and most cost-efficient molecules to test is one reason new medicines take so long to develop, as well as a key driver of high prescription drug prices.
To help scientists make cost-aware choices, MIT researchers developed an algorithmic framework to automatically identify optimal molecular candidates, which minimizes synthetic cost while maximizing the likelihood candidates have desired properties. The algorithm also identifies the materials and experimental steps needed to synthesize these molecules. Learn more

Explainable AI for Rational Antibiotic Discovery
Researchers now employ artificial intelligence (AI) models based on deep learning to make functional predictions about big datasets. While the concepts behind these networks are well established, their inner workings are often invisible to the user. The emerging area of explainable AI (xAI) provides model interpretation techniques that empower life science researchers to uncover the underlying basis on which AI models make such predictions.In this month’s episode, Deanna MacNeil from The Scientist spoke with Jim Collins from the Massachusetts Institute of Technology to learn how researchers are using explainable AI and artificial neural networks to gain mechanistic insights for large scale antibiotic discovery. Learn more
When an antibiotic fails: MIT scientists are using AI to target “sleeper” bacteria
Since the 1970s, modern antibiotic discovery has been experiencing a lull. Now the World Health Organization has declared the antimicrobial resistance crisis as one of the top 10 global public health threats.When an infection is treated repeatedly, clinicians run the risk of bacteria becoming resistant to the antibiotics. But why would an infection return after proper antibiotic treatment? One well-documented possibility is that the bacteria are becoming metabolically inert, escaping detection of traditional antibiotics that only respond to metabolic activity. When the danger has passed, the bacteria return to life and the infection reappears.
“Resistance is happening more over time, and recurring infections are due to this dormancy,” says Jackie Valeri, a former MIT-Takeda Fellow (centered within the MIT Abdul Latif Jameel Clinic for Machine Learning in Health) who recently earned her PhD in biological engineering from the Collins Lab. Valeri is the first author of a new paper published in this month’s print issue of Cell Chemical Biology that demonstrates how machine learning could help screen compounds that are lethal to dormant bacteria. Learn more

A new computational technique could make it easier to engineer useful proteins
To engineer proteins with useful functions, researchers usually begin with a natural protein that has a desirable function, such as emitting fluorescent light, and put it through many rounds of random mutation that eventually generate an optimized version of the protein.This process has yielded optimized versions of many important proteins, including green fluorescent protein (GFP). However, for other proteins, it has proven difficult to generate an optimized version. MIT researchers have now developed a computational approach that makes it easier to predict mutations that will lead to better proteins, based on a relatively small amount of data.
Using this model, the researchers generated proteins with mutations that were predicted to lead to improved versions of GFP and a protein from adeno-associated virus (AAV), which is used to deliver DNA for gene therapy. They hope it could also be used to develop additional tools for neuroscience research and medical applications.
“Protein design is a hard problem because the mapping from DNA sequence to protein structure and function is really complex. There might be a great protein 10 changes away in the sequence, but each intermediate change might correspond to a totally nonfunctional protein. It’s like trying to find your way to the river basin in a mountain range, when there are craggy peaks along the way that block your view. The current work tries to make the riverbed easier to find,” says Ila Fiete, a professor of brain and cognitive sciences at MIT, a member of MIT’s McGovern Institute for Brain Research, director of the K. Lisa Yang Integrative Computational Neuroscience Center, and one of the senior authors of the study.
Regina Barzilay, the School of Engineering Distinguished Professor for AI and Health at MIT, and Tommi Jaakkola, the Thomas Siebel Professor of Electrical Engineering and Computer Science at MIT, are also senior authors of an open-access paper on the work, which will be presented at the International Conference on Learning Representations in May. MIT graduate students Andrew Kirjner and Jason Yim are the lead authors of the study. Other authors include Shahar Bracha, an MIT postdoc, and Raman Samusevich, a graduate student at Czech Technical University. Learn more
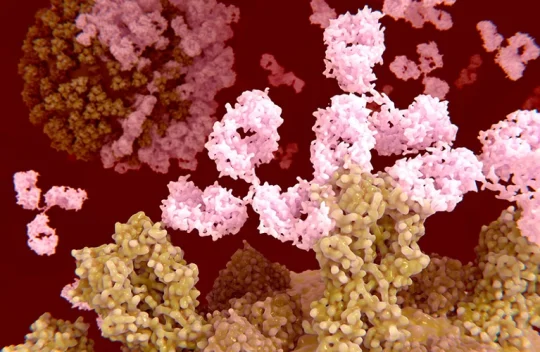
‘A landmark moment’: scientists use AI to design antibodies from scratch
Researchers have used generative artificial intelligence (AI) to help them make completely new antibodies for the first time.The proof-of-principle work, reported this week in a preprint on bioRxiv, raises the possibility of bringing AI-guided protein design to the therapeutic antibody market, which is worth hundreds of billions of dollars.
Antibodies — immune molecules that strongly attach to proteins implicated in disease — have conventionally been made using brute-force approaches that involve immunizing animals or screening vast numbers of molecules.
AI tools that can shortcut those costly efforts have the potential to “democratize the ability to design antibodies”, says study co-author Nathaniel Bennett, a computational biochemist at the University of Washington in Seattle. “Ten years from now, this is how we’re going to be designing antibodies.”
“It’s a really promising piece of research” that represents an important step in applying AI protein-design tools to making new antibodies, says Charlotte Deane, an immuno-informatician at the University of Oxford, UK. Learn more

Jim Collins: Discovery of the First New Structural Class of Antibiotics in Decades, Using A.I.
Jim Collins is one of the leading biomedical engineers in the world. He’s been elected to all 3 National Academies (Engineering, Science, and Medicine) and is one of the founders of the field of synthetic biology. In this conversation, we reviewed the seminal discoveries that he and his colleagues are making at the Antibiotics-AI Project at MIT. Learn more
Why Casey Left Substack, Elon Musk and Drugs, and an A.I. Antibiotic Discovery
Casey is taking his newsletter Platformer off Substack, as criticism over the company’s handling of pro-Nazi content grows. Then, The Wall Street Journal spoke with witnesses who said that Elon Musk had used LSD, cocaine, ecstasy and psychedelic mushrooms, worrying some directors and board members of his companies. And finally, how researchers found a new class of antibiotics with the help of an artificial intelligence algorithm used to win the board game Go.Today’s guests:
Kirsten Grind, enterprise reporter for The Wall Street Journal
Felix Wong, postdoctoral fellow at M.I.T. and co-founder of Integrated Biosciences
Additional Reading:
Why Platformer is leaving Substack.
Elon Musk has reportedly used illegal drugs, worrying leaders at Tesla and SpaceX.
Researchers have discovered a new class of antibiotics using A.I. Learn more




