Tag: Gabriele Corso

Meet the newest Forbes 30 under 30
Gabriele Corso was a computer science PhD student at MIT's Computer Science and Artificial Intelligence Laboratory (CSAIL), where his research focused on the intersection of machine learning and molecular biology. His cofounder Saro Passaro was also a research scientist at MIT and previously worked at Meta. The two have trained AI models for predicting biomolecular structures and how molecules interact within the body, which could eventually help with drug discovery. Used by thousands of global organizations and downloaded more than 1M times, these open source models are the basis for Boltz, a company Corso and Passaro cofounded with Jeremy Wohlwend to improve therapeutic design using AI. Learn more
What will be the first AI-designed drug? These disease-fighting antibodies are top contenders
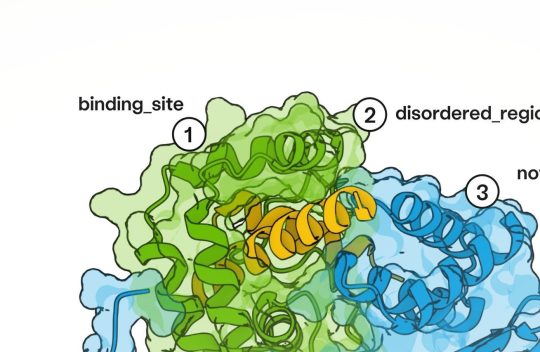
Antibodies — immune proteins that recognize foreign molecules, such as those made by pathogens, with exquisite specificity — have been a challenge for AI to design. AI models such as AlphaFold have struggled to predict the shape of flexible loop regions of antibodies, which they use to recognize their targets.But new tools developed in the past year — including an updated version of AlphaFold — have proved better at modelling these flexible regions, says Gabriele Corso, a machine-learning scientist at the Massachusetts Institute of Technology in Cambridge. Progress in antibody design has followed.
In October, Corso and his colleagues described the BoltzGen model in a preprint, showing that it can adroitly design ‘nanobodies’ — small, simple antibodies resembling molecules made by sharks and camels — against proteins implicated in cancer, viral and bacterial infections and other diseases. In most cases, the researchers identified antibodies with strong target binding after expressing just 15 of the most-promising designs in cells and testing them in laboratory experiments. However, the molecules were not tested in disease models. Learn more
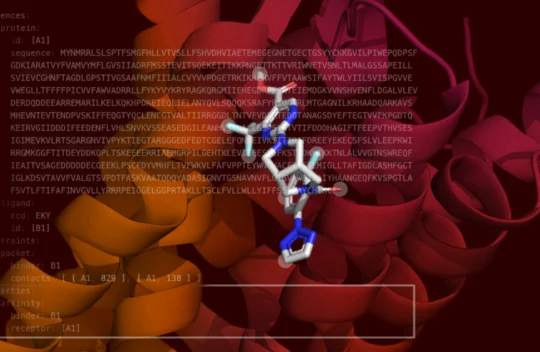
Boltz-2 Released to Democratize AI Molecular Modeling for Drug Discovery
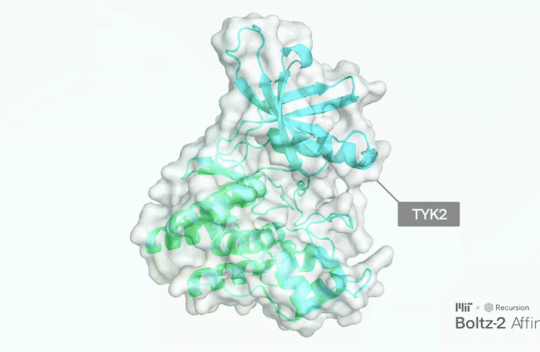
Researchers from the Massachusetts Institute of Technology (MIT) Jameel Clinic for Machine Learning in Health have announced the open-source release of Boltz-2, which now predicts molecular binding affinity at newfound speed and accuracy to democratize commercial drug discovery. The model is available under the highly permissive MIT license, which allows commercial drug developers to use the model internally and apply their own proprietary data. Learn more
The Prototype: This AI Model Could Make It Faster To Find New Medicines
The 2024 Nobel Prize in Chemistry was awarded in part to Deepmind’s Demis Hassabis and John Jumper for the development of AlphaFold–an AI model that predicts the structure of proteins, the complex chemicals essential to making our bodies work. Since its inception, this model and others like it have been put to use in laboratories around the world, enabling new biological discoveries.Now a team from MIT and pharmaceutical company Recursion, with support from Cancer Grand Challenges, have developed a tool that takes these principles further–and may help researchers find new medicines more quickly. Called Boltz-2, this open-source generative AI model can not only predict the structure of proteins, it can also predict its binding affinity–that is, how well a potential drug is able to interact with that protein. This is crucial in the early stages of developing a new medicine. Learn more

MIT researchers introduce Boltz-1, a fully open-source model for predicting biomolecular structures
MIT scientists have released a powerful, open-source AI model, called Boltz-1, that could significantly accelerate biomedical research and drug development. Developed by a team of researchers in the MIT Jameel Clinic for Machine Learning in Health, Boltz-1 is the first fully open-source model that achieves state-of-the-art performance at the level of AlphaFold3, the model from Google DeepMind that predicts the 3D structures of proteins and other biological molecules. MIT graduate students Jeremy Wohlwend and Gabriele Corso were the lead developers of Boltz-1, along with MIT Jameel Clinic Research Affiliate Saro Passaro and MIT professors of electrical engineering and computer science Regina Barzilay and Tommi Jaakkola. Wohlwend and Corso presented the model at a Dec. 5 event at MIT’s Stata Center, where they said their ultimate goal is to foster global collaboration, accelerate discoveries, and provide a robust platform for advancing biomolecular modeling. Learn more