Tag: drug discovery

One Survivor’s AI Breakthrough Predicts Cancer Years Ahead
AI Decoded focusses on one of the most urgent, tangible uses of artificial intelligence: health care — we speak to Dr Regina Barzilay, an MIT professor who is building machine-learning AI models to predict disease. She herself was diagnosed with breast cancer in 2014, and has used that experience and knowledge to target her research towards prevention — the AI model she and her team built, named MIRAI, is now able to detect a patient’s risk of developing breast cancer within five years. Are we on the brink of a revolution in treating cancer for everyone? Find out on AI Decoded... Joining presenter Christian Fraser is AI Decoded co-host Stephanie Hare and the BBC's AI correspondent Marc Cieslak Learn more
Meet the newest Forbes 30 under 30
Gabriele Corso was a computer science PhD student at MIT's Computer Science and Artificial Intelligence Laboratory (CSAIL), where his research focused on the intersection of machine learning and molecular biology. His cofounder Saro Passaro was also a research scientist at MIT and previously worked at Meta. The two have trained AI models for predicting biomolecular structures and how molecules interact within the body, which could eventually help with drug discovery. Used by thousands of global organizations and downloaded more than 1M times, these open source models are the basis for Boltz, a company Corso and Passaro cofounded with Jeremy Wohlwend to improve therapeutic design using AI. Learn more
What will be the first AI-designed drug? These disease-fighting antibodies are top contenders
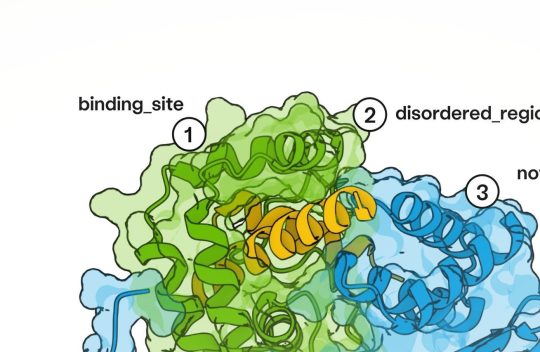
Antibodies — immune proteins that recognize foreign molecules, such as those made by pathogens, with exquisite specificity — have been a challenge for AI to design. AI models such as AlphaFold have struggled to predict the shape of flexible loop regions of antibodies, which they use to recognize their targets.But new tools developed in the past year — including an updated version of AlphaFold — have proved better at modelling these flexible regions, says Gabriele Corso, a machine-learning scientist at the Massachusetts Institute of Technology in Cambridge. Progress in antibody design has followed.
In October, Corso and his colleagues described the BoltzGen model in a preprint, showing that it can adroitly design ‘nanobodies’ — small, simple antibodies resembling molecules made by sharks and camels — against proteins implicated in cancer, viral and bacterial infections and other diseases. In most cases, the researchers identified antibodies with strong target binding after expressing just 15 of the most-promising designs in cells and testing them in laboratory experiments. However, the molecules were not tested in disease models. Learn more
MIT scientists debut a generative AI model that could create molecules addressing hard-to-treat diseases
More than 300 people across academia and industry spilled into an auditorium to attend a BoltzGen seminar on Thursday, Oct. 30, hosted by the Abdul Latif Jameel Clinic for Machine Learning in Health (MIT Jameel Clinic). Headlining the event was MIT PhD student and BoltzGen’s first author Hannes Stärk, who had announced BoltzGen just a few days prior. Learn more
2025’s Fiercest Women in Life Sciences
The life sciences mission of Najat Khan, Ph.D., was forged in hospital hallways.Her parents, a trauma surgeon and a gynecologist-turned-radiologist, “didn't believe in babysitters,” she joked in an interview. The extensive time she therefore spent in hospitals throughout her childhood showed Khan the life-changing impact of innovative medicines and set her on the path to the C-suites of Johnson & Johnson and Recursion Pharmaceuticals. Learn more
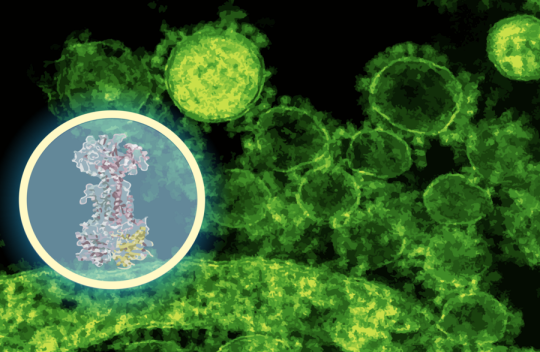
AI maps how a new antibiotic targets gut bacteria
“This discovery speaks to a central challenge in antibiotic development,” says Jon Stokes, senior author of a new paper on the work, assistant professor of biochemistry and biomedical sciences at McMaster, and research affiliate at MIT’s Abdul Latif Jameel Clinic for Machine Learning in Health. “The problem isn’t finding molecules that kill bacteria in a dish — we’ve been able to do that for a long time. A major hurdle is figuring out what those molecules actually do inside bacteria. Without that detailed understanding, you can’t develop these early-stage antibiotics into safe and effective therapies for patients.” Learn more